Coronary artery disease is the most common heart disease in the US, killing over 370 000 people annually. [1] Many factors contribute to the development of coronary artery disease and early detection of symptoms can greatly reduce the risk of heart infarcts and death. In recent years, optical coherence tomography (OCT) has become increasingly useful in the study of coronary artery tissues and their characterization. OCT is a catheter-based imaging modality that uses infrared light to acquire cross sectional images of the coronary artery (Figure 1). Compared to other non-invasive modalities like MRI or CT, OCT provides information on the tissue contents (e.g. its layered structure) of the artery, with a resolution that exceeds modalities such as intravascular ultrasound (IVUS). [2]
An article titled “Characterization of coronary artery pathological formations from OCT imaging using deep learning” was published in the biomedical optics express journal in 2018 by leading author Atefeh Abdolmanafi, who was a PhD candidate at the time. Under the supervision of Luc Duong from the department of software and IT engineering of École de technologie supérieure, Atefeh’s goal was to develop a deep learning model to characterize different coronary artery tissues and different pathological formations that can occur prior to the development of coronary artery disease. The main interest in having such a model is to enable doctors to detect signs of coronary artery disease earlier and more reliably.
I met with Atefeh in early March 2019 for a Q&A to learn more about the goals of her work, the model she developed, and her vision for applying deep learning approaches in the medical field.

How did this project about coronary artery disease come about?
It was proposed by my supervisor Luc. I had a background in physics, so he thought it could be a good project for me since I would also understand the physics behind the OCT system. At the beginning, I started by understanding the OCT system. Then I talked to a doctor in Ste-Justine hospital and he explained what lesions they were looking for in the images and how important it was for them to make the detection automatic. In some cases, even for the doctors it is challenging to differentiate normal tissues from the lesions. They needed an automatic model to help them with that. We decided to apply machine learning.
Basically, we started with OCT images of normal coronary arteries to detect their three-layer structure. The first two layers, intima and media, were the most important ones to analyze, so we wanted to be able to segment them first. Then, we extended the model to detect pathological lesions that indicate coronary artery disease. I specifically worked on Kawasaki disease which is a disease in children.
You talk a lot about Kawasaki disease (KD) in the article. Can you elaborate a bit more on what dangers it presents for children?
It is a disease in infants, and the reason for it is unknown yet but it can lead to coronary artery disease if not treated. If doctors can diagnose KD early enough and if the children have a good treatment, they do not suffer from heart disease later in life. So, it is important for doctors to know what is the impact on coronary arteries at the beginning of the disease. Sometimes the first symptoms are just the thickening of the layers and no lesions are observed, that is called intima thickening. It was important to detect the intima and media first just to measure how thick they are.
Because that is one of the first signs of the disease?
Yes, this is the first symptom that can be detected and after that the other problems like calcification, vascularisation or macrophage accumulation can occur. However, my focus was not on KD in the beginning, I wanted to develop something that could be extended to any coronary artery disease. That’s why when we were developing the model, we were thinking about other cases also, not just KD. Now I am working with an adult cardiologist to extend the model to adult cardiology.
You mention how much data was available to train the model. It is known that for deep learning applications, the more data you have, the better it is. How did you deal with the fact that you didn’t have that much data?
It is true that in medical imaging the problem is that we don’t have access to a lot of data. For this paper I had data from 45 patients in total, but I couldn’t use all the data because of the quality of the acquisitions. I had enough data, but I had to manage how to train deep convolutional neural network (CNN) models with this data. In practice we need thousands of images to train a model, so the first thing I thought about was to use pretrained networks so that I wouldn’t need to initialize the weights of the model with something random, I could initialize my model with the weights of the pretrained network. Although these pretrained networks are not meant for medical images, we could still use their learned weights instead of starting from scratch. It helped me a lot to accelerate learning and have a good model.
Other than the limited amount of data, did you encounter any other difficulties in training, such as hardware limitations?
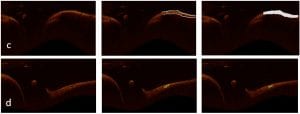
We had a very powerful computer with a good GPU, so I did not encounter that problem. The only problem was that I needed to decide which loss function and optimizer to use. For the CNN models I used cross-entropy for the loss function and stochastic gradient descent as the optimizer, that gave me a good precision. After that, when I wanted to train the fully convolutional networks, I encountered the problem of class imbalance since in some cases the images have a lot of background and a very small region of interest to segment (Figure 2d).
Why did you use the majority voting approach to determine the final classification of the tissues?
In my first paper working with coronary arteries, I only wanted to detect the intima and media. In that paper I used a pretrained CNN called AlexNet and after training on medical images I got very high accuracy for the classification of the first 2 layers. When I wanted to extend the network to classify more tissues like pathological formations, I could not get good results. Doctors need models with high precision or else they’re better off classifying the tissue on their own. So my model with around 70% accuracy was useless and I had to try other networks.
AlexNet is a shallow network, so it can give me abstract information about the borders and a little bit of textural information. VGG-19 is a deeper network so it can give me more information about the textures in the image. The size of the filters is different, so it gives different information about the image. Complex networks like inception V3 also gave me more detailed information. I could get a lot of information using the 3 networks together. Each network worked well for a certain type of tissue. AlexNet worked better to detect intima, VGG-19 for lesions, and Inception V3 worked well in cases where lesions were very small. So, I thought that maybe if I can use them as feature extractors, and then use random forest to do the classification on all the features the networks generate, I could get a better result and reduce training time.
Why did you use Random forest for the final classification?
In my first paper, where I only wanted to segment intima and media layers, I started with AlexNet and I wanted to see if it is better to use it as a classifier or a feature extractor. The other classifiers I used were SVM and random forest and I compared them to the classification of AlexNet. I found that it was better to use AlexNet as the feature extractor and then use Random forest on those features to make the final classification. I came to the same conclusion after testing other networks as classifiers, so I kept this strategy in my later work.
You mentioned earlier that you were working on adapting the network for adults instead of children. What are the additional steps it takes to do this adaptation? Does the network need to be changed?
We don’t know exactly since we haven’t started working on this yet. We are collecting data right now, but I don’t think we need to change anything in the network. We don’t even need to retrain the network since I use it as a feature extractor. I think we only need to retrain the classifier at the end of the network. We can add as many lesions as we want to detect, we only need to decide with the doctors which lesions are important to detect in adults that are not present in children. Once that is established it should be easy to extend the model for more lesions. We will see… I am 90% confident this should work, however there is always this 10% chance that, once we do the experiments, it doesn’t. In that case we will probably need to change the model a little bit, but I hope not!

In your opinion, how long will it take before approaches like the one you developed could be really used in a clinical setting?
We are currently working on it and since the publication of the paper we are discussing, we have made improvements to the model. I can now look at a complete sequence of images from an acquisition and decide which images are normal and which ones seem problematic. For the normal images the model extracts the 2 important layers, and for the problematic images the model looks for lesions and then characterizes them. I think it is quite useful for doctors right now. We are talking with companies to possibly add the model to their system directly. The programming part needs to be optimized by software engineers, but after that it should be ready to be used clinically.
Are doctors reluctant to use this type of software?
Not at all! The doctor I am working with is very interested. He explained to me that he needs such a model to help the doctors. It is not going to replace the doctors for sure, but it can help them a lot, so they don’t have to spend a lot of time or analyzing the images one by one.
A current problem in deep learning applied to the medical field is that sometimes doctors hesitate to let models like this to even help them.
Yes, I am personally against replacing the humans by artificial intelligence. I think that it should just help them where they most need the help. I don’t think we can trust these models 100%, we need an expert behind it to double-check. But they can help doctors. During the development of my model the doctors at Ste-Justine were very motivated and they always found availabilities to meet me and suggest improvements to the model. I guess I am very lucky!
Currently Atefeh is doing her post doctorate at the LIV4D laboratory in Polytechnique Montréal.
____________________________________________________________________________________________________
Reference:
[1] CDC, NCHS. Underlying Cause of Death 1999-2013 on CDC WONDER Online Database, released 2015. Data are from the Multiple Cause of Death Files, 1999-2013, as compiled from data provided by the 57 vital statistics jurisdictions through the Vital Statistics Cooperative Program.
[2] Atefeh Abdolmanafi, Luc Duong, Nagib Dahdah, Ibrahim Ragui Adib, and Farida Cheriet, “Characterization of coronary artery pathological formations from OCT imaging using deep learning,” Biomed. Opt. Express 9, 4936-4960 (2018)